Research in the Grillo lab is aimed at understanding the mechanisms of adaptation. We utilize a broad range of evolutionary genetics and evolutionary ecology approaches that involve field, lab, and greenhouse experiments. Below is information on the main study systems and projects currently underway, as well as lab publications.
Genetic basis of adaptation in the wild relatives of rice
Understanding the genetic basis of adaptation has been a seminal goal in evolutionary biology for over a century. The development of quantitative trait loci (QTL) mapping has provided an empirical means for testing theories of the genetic architecture of adaptation. However, QTL regions can be quite large and may encompass hundreds of genes. To fully understand the genetic basis of adaptation we need to move beyond QTL and identify the causal genes and mutations.
The wild progenitors of cultivated rice, Oryza nivara and Oryza rufipogon, present an ideal system for studying the mechanisms of adaptation. Oryza nivara and O. rufipogon are recently diverged sister species that have undergone marked adaptive differentiation to distinct habitats in the tropics and sub-tropics of Asia. O. nivara is a highly self-fertilized annual that lives in vernal pools that desiccate during the dry season. O. rufipogon is an outcrossing perennial that inhabits wetlands such as marshes and river banks. Current projects in the lab are aimed at understanding the genetic basis of mating system and life history evolution in this system. Here we are conducting QTL mapping and fine mapping approaches to further dissect QTL regions to find the causal genetic elements controlling adaptive trait variation. I am also interested in studying parallel evolution of closely related species in the new world tropics (O. glumaepatula) and other patterns of evolution in the Oryza genus.
The wild progenitors of cultivated rice, Oryza nivara and Oryza rufipogon, present an ideal system for studying the mechanisms of adaptation. Oryza nivara and O. rufipogon are recently diverged sister species that have undergone marked adaptive differentiation to distinct habitats in the tropics and sub-tropics of Asia. O. nivara is a highly self-fertilized annual that lives in vernal pools that desiccate during the dry season. O. rufipogon is an outcrossing perennial that inhabits wetlands such as marshes and river banks. Current projects in the lab are aimed at understanding the genetic basis of mating system and life history evolution in this system. Here we are conducting QTL mapping and fine mapping approaches to further dissect QTL regions to find the causal genetic elements controlling adaptive trait variation. I am also interested in studying parallel evolution of closely related species in the new world tropics (O. glumaepatula) and other patterns of evolution in the Oryza genus.
Examples of O. nivara and O. rufipogon habitats. This is a wetland that Mike visited in Thailand. O. rufipogon inhabits the deeper wetland habitats that maintain water levels throughout the year. O. nivara lives in shallow wetlands that desiccate during the dry season. Only O. rufipogon was found at this site, but the two species can be found in sympatry. Distinct flowering time is likely a key isolating barrier between these species.
Genetics of plant-microbe interactions in legumes
Mutualisms are ubiquitous in nature, contributing to numerous ecological processes and playing profound roles in ecosystem functioning. A striking feature of plant-microbe mutualisms is the high degree of variation maintained for both partner choice (i.e. the symbiotic partners a plant forms associations with) and partner quality (the fitness benefits a plant receives from a particular symbiont). The mutualism between plants in the legume family (Fabaceae) and rhizobia (nitrogen-fixing bacteria) is one of the most economically and ecologically important interactions on the planet. Currently my lab is conducting genome wide association studies (GWAS) to identify the genetic basis of mutualism variation (e.g. partner choice) in the model legume Medicago truncatula, and its symbiotic rhizobia (Ensifer spp.). We are also conducting population genomic studies in Ensifer collected from the native range in Europe.
Beyond classic microbial mutualists (e.g. rhizobia and AMF) plants intimately associate with a remarkable diversity of bacterial and fungal species (i.e. the microbiome); and these associations can influence plant traits, distributions, and fitness. Only with modern meta-genomic sequencing approaches are we beginning to thoroughly characterize these communities and understand the factors that structure them. We are conducting various experiments to examine the genetic and environmental drivers that govern legume associated microbiomes.
Beyond classic microbial mutualists (e.g. rhizobia and AMF) plants intimately associate with a remarkable diversity of bacterial and fungal species (i.e. the microbiome); and these associations can influence plant traits, distributions, and fitness. Only with modern meta-genomic sequencing approaches are we beginning to thoroughly characterize these communities and understand the factors that structure them. We are conducting various experiments to examine the genetic and environmental drivers that govern legume associated microbiomes.
Medicago truncatula population on the coast in Corsica that Mike collected from. The Medicago plants are the green clover like plant growing among the rocks. Mike cultured hundreds of rhizobia from the native range in Europe. We are using these isolates for numerous experiments in the lab.
Coevolutionary diversification in astragalus
Coevolution has been considered a primary driver of diversification since the origins of evolutionary thought. Surprisingly, limited experimental evidence exists directly testing this hypothesis and elucidating the mechanisms of coevolutionary diversification – particularly at the microevolutionary level. The central challenge in this pursuit is connecting coevolution to a form of reproductive isolation. We are testing for coevolutionary diversification in Astragalus, the single most species-rich genus of flowering plants (ca. 3,200 spp.). The genus is distributed mainly in arid to semiarid temperate regions and is particularly diverse in western North America (400+ spp.). Astragalus is renowned for specialized interactions with both mutualistic (e.g. rhizobia and fungal endophytes) and antagonistic (e.g. herbivorous insects) interactors. We are conducting a broad range of experiments to examine the role of coevolutionary interactions in contributing to the extraordinary diversity in this genus. We focus primarily on A. lentiginosus the most taxonomically diverse species in the North American flora with over 40 recognized varieties.
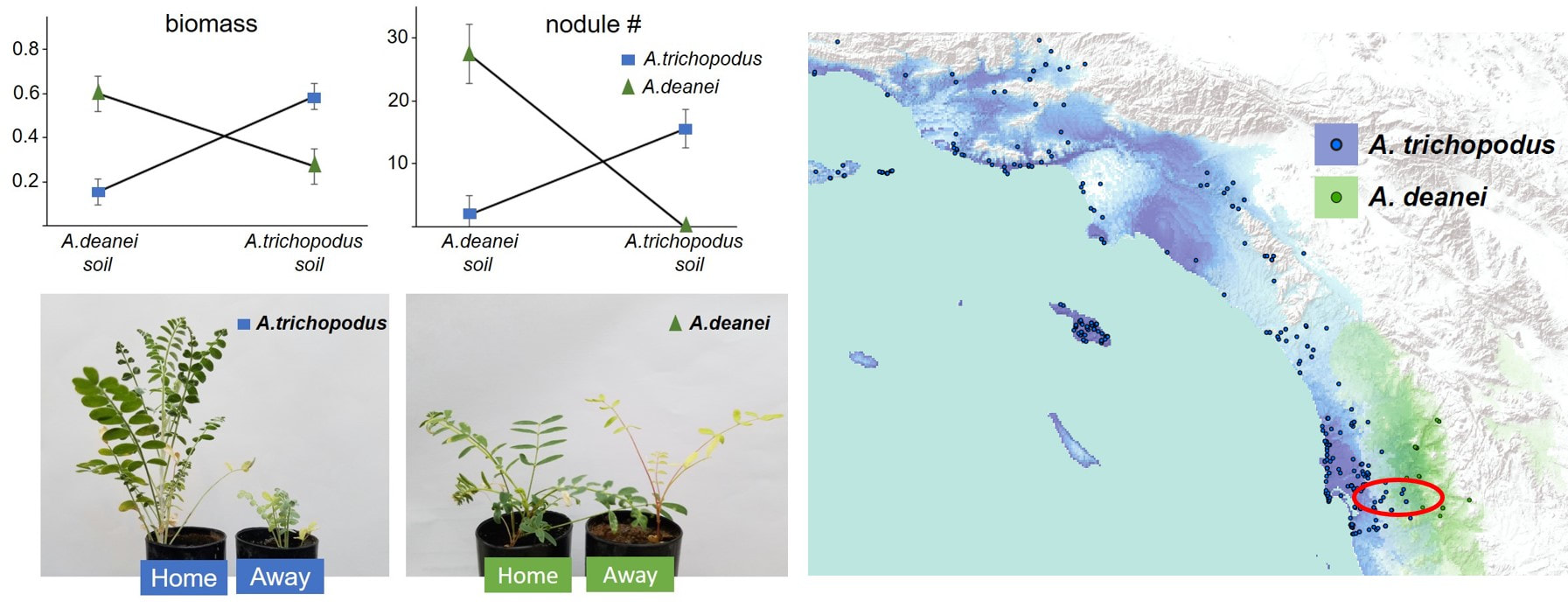
We hypothesize that interactions with rhizobia mediate local adaptation of Astragalus to soil habitats and contribute to ecogeographic reproductive isolation. On the left are results of a reciprocal transplant experiment where we grew plants on their home and away soil demonstrating fitness tradeoffs. Both species performed better on their home soil. On the right are species distribution models showing their distributions. The red circle represents where the plants for the experiment were collected. Even those these species were collected less than 10 miles from each other, they are adapted to their own soil and rhizobia.
lab publications
Submitted manuscripts:
del-Canto, A., Sanz-Saez, A., Heath, K., Grillo, M., Heras, J., Lacuesta, M. Agrochemical inputs and management have a greater impact on common bean rhizobia diversity than drought. In review.
Kilbane, E., Crews, T., DeHaan, L., Ohsowski, B., Grillo, M., Dybzinski, R. Modeling carbon allocation strategies for high-yielding perennial crops. In revision.
Peer reviewed publications:
Batstone RT, Lindgren H, Allsup CM, Goralka LA, Riley AB, Grillo MA, Marshall-Colon A, Heath KD. Genome-Wide Association Studies across Environmental and Genetic Contexts Reveal Complex Genetic Architecture of Symbiotic Extended Phenotypes. mBio. 2022 Dec 20;13(6):e0182322. doi: 10.1128/mbio.01823-22.
Riley AB, Grillo MA, Epstein B, Tiffin P, Heath KD. (2022) Discordant population structure among rhizobium divided genomes and their legume hosts. Molecular Ecology. Sep 26. doi: 10.1111/mec.16704.
Gutierrez, A., and Grillo, M.A. (2022) Effects of Domestication on Plant–Microbiome Interactions. Plant and Cell Physiology pcac108, https://doi.org/10.1093/pcp/pcac108
Epstein B, Burghardt LT, Heath KD, Grillo MA, Kostanecki A, Hämälä T, Young ND, Tiffin P. (2022) Combining GWAS and population genomic analyses to characterize coevolution in a legume-rhizobia symbiosis. Molecular Ecology. Jul 6. doi: 10.1111/mec.16602.
*Brown, S.P., *Grillo, M.A., Podowski, J.C., Heath, K.D. (2020), Soil origin and plant genotype structure distinct microbiome compartments in the model legume Medicago truncatula. Microbiome 8, 139. *co-first authors decided by coin flip
Grillo M.A., S. de Mita, P.V. Burke, K.D. Heath. (2016), Intra-population genomics in a model mutualist: population structure and candidate symbiosis genes under selection in Medicago truncatula. Evolution 70: 2704-2717.
Grillo M.A., J.R. Stinchcombe, K.D. Heath. (2016), Nitrogen addition does not influence pre-infection partner choice in the legume-rhizobium symbiosis. American Journal of Botany 103: 1763-1770.
Heath, K. D. and Grillo, M. A. (2016), Rhizobia: tractable models for bacterial evolutionary ecology. Environmental Microbiology 18: 4307-4311.
Grillo, M.A., M. Hammond, C. Li, L. Wang, D.W. Schemske. (2013), Genetic architecture of flowering time differentiation between locally adapted populations of Arabidopsis thaliana. New Phytologist 197: 1321-1331.
Grillo, M.A, C. Li, A.M. Fowlkes, T.M. Briggeman, A. Zhou, D.W. Schemske, and T. Sang. (2009), Genetic architecture for the adaptive origin of annual wild rice, Oryza nivara. Evolution 63: 870-883 .
Zhao, Q-F., G. Wang, Q-X Li, S-R Ma,Y. Cui, M.A. Grillo. (2006), Genetic diversity of five Kobresia species along the eastern Qinghai-Tibet plateau in China. Hereditas 143: 34-41.
del-Canto, A., Sanz-Saez, A., Heath, K., Grillo, M., Heras, J., Lacuesta, M. Agrochemical inputs and management have a greater impact on common bean rhizobia diversity than drought. In review.
Kilbane, E., Crews, T., DeHaan, L., Ohsowski, B., Grillo, M., Dybzinski, R. Modeling carbon allocation strategies for high-yielding perennial crops. In revision.
Peer reviewed publications:
Batstone RT, Lindgren H, Allsup CM, Goralka LA, Riley AB, Grillo MA, Marshall-Colon A, Heath KD. Genome-Wide Association Studies across Environmental and Genetic Contexts Reveal Complex Genetic Architecture of Symbiotic Extended Phenotypes. mBio. 2022 Dec 20;13(6):e0182322. doi: 10.1128/mbio.01823-22.
Riley AB, Grillo MA, Epstein B, Tiffin P, Heath KD. (2022) Discordant population structure among rhizobium divided genomes and their legume hosts. Molecular Ecology. Sep 26. doi: 10.1111/mec.16704.
Gutierrez, A., and Grillo, M.A. (2022) Effects of Domestication on Plant–Microbiome Interactions. Plant and Cell Physiology pcac108, https://doi.org/10.1093/pcp/pcac108
Epstein B, Burghardt LT, Heath KD, Grillo MA, Kostanecki A, Hämälä T, Young ND, Tiffin P. (2022) Combining GWAS and population genomic analyses to characterize coevolution in a legume-rhizobia symbiosis. Molecular Ecology. Jul 6. doi: 10.1111/mec.16602.
*Brown, S.P., *Grillo, M.A., Podowski, J.C., Heath, K.D. (2020), Soil origin and plant genotype structure distinct microbiome compartments in the model legume Medicago truncatula. Microbiome 8, 139. *co-first authors decided by coin flip
Grillo M.A., S. de Mita, P.V. Burke, K.D. Heath. (2016), Intra-population genomics in a model mutualist: population structure and candidate symbiosis genes under selection in Medicago truncatula. Evolution 70: 2704-2717.
Grillo M.A., J.R. Stinchcombe, K.D. Heath. (2016), Nitrogen addition does not influence pre-infection partner choice in the legume-rhizobium symbiosis. American Journal of Botany 103: 1763-1770.
Heath, K. D. and Grillo, M. A. (2016), Rhizobia: tractable models for bacterial evolutionary ecology. Environmental Microbiology 18: 4307-4311.
Grillo, M.A., M. Hammond, C. Li, L. Wang, D.W. Schemske. (2013), Genetic architecture of flowering time differentiation between locally adapted populations of Arabidopsis thaliana. New Phytologist 197: 1321-1331.
Grillo, M.A, C. Li, A.M. Fowlkes, T.M. Briggeman, A. Zhou, D.W. Schemske, and T. Sang. (2009), Genetic architecture for the adaptive origin of annual wild rice, Oryza nivara. Evolution 63: 870-883 .
Zhao, Q-F., G. Wang, Q-X Li, S-R Ma,Y. Cui, M.A. Grillo. (2006), Genetic diversity of five Kobresia species along the eastern Qinghai-Tibet plateau in China. Hereditas 143: 34-41.